About voluba-mriwarp
Note: voluba-mriwarp is still in development. You may still encounter bugs when using it.
voluba (Volumetric Brain Anchoring) offers tools to connect volumetric imaging data to multilevel atlases and in this way makes it accessible for analysis with the siibra toolsuite.
voluba-mriwarp is a desktop application that integrates whole-brain T1-weighted MRI scans into the anatomical context of the EBRAINS Human Brain Atlas. It incorporates all required components like skull stripping, registration to MNI ICBM 152 2009c Nonlinear Asymmetric space, and detailed analysis with the siibra toolsuite. The corresponding functionalities are provided via open-source tools like HD-BET1, ANTs and siibra-python. voluba-mriwarp is primarily designed for Windows 10 but can also be executed on Linux.
Your data remains on your computer
We designed voluba-mriwarp as a desktop application. Therefore, the input MRI dataset will not be uploaded to any online services. It remains only on your local computer.
Warping brain data to a standardized space like MNI152 enables anchoring of whole-brain MRI scans to atlas volumes and therefore facilitates analysis in a detailed anatomical context. However, reasonable registration requires various steps that need optimization effort. voluba-mriwarp aims to simplify this workflow. With this application, you avoid installing multiple tools and tweaking parameters for a proper registration result. Instead, voluba-mriwarp is an easy-to-install and easy-to-use tool combining all necessary steps into one pipeline.
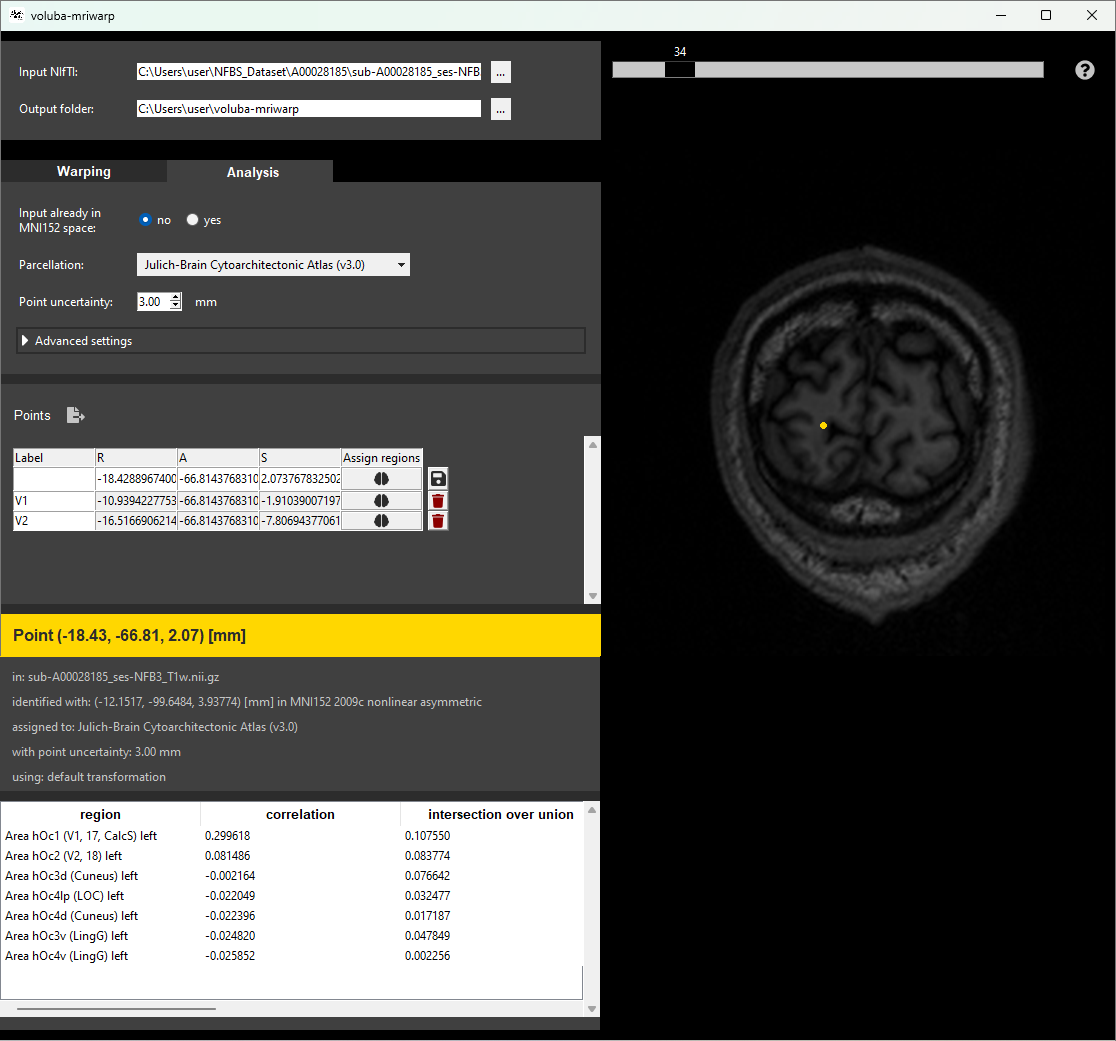
voluba-mriwarp applies a set of predefined parameters to remove the skull and warp the input brain scan to MNI152 space. You can utilize the warping results in voluba-mriwarp to interactively analyze points by making a probabilistic assignment of coordinates in the input space to brain regions of the EBRAINS Human Brain Atlas. To get an overview of more information about a brain region, you can access siibra-explorer through the application. For further analysis, voluba-mriwarp offers to export the anatomical assignments together with linked multimodal data features like receptor densities, cell distributions or brain connectivity.
-
Isensee F, Schell M, Tursunova I, Brugnara G, Bonekamp D, Neuberger U, Wick A, Schlemmer HP, Heiland S, Wick W, Bendszus M, Maier-Hein KH, Kickingereder P. Automated brain extraction of multi-sequence MRI using artificial neural networks. Hum Brain Mapp. 2019; 1–13. https://doi.org/10.1002/hbm.24750 ↩